[1] Huang, Xiaowan, et al. "Nanoreceptors promote mutant p53 protein degradation by mimicking selective autophagy receptors." Nature Nanotechnology (2024): 1-9.
[2] Rajbanshi B, Guruacharya A, Mandell J W, et al. Localization, induction, and cellular effects of tau phosphorylated at threonine 217 1[J]. Alzheimer's & Dementia, 2023, 19(7): 2874-2887.
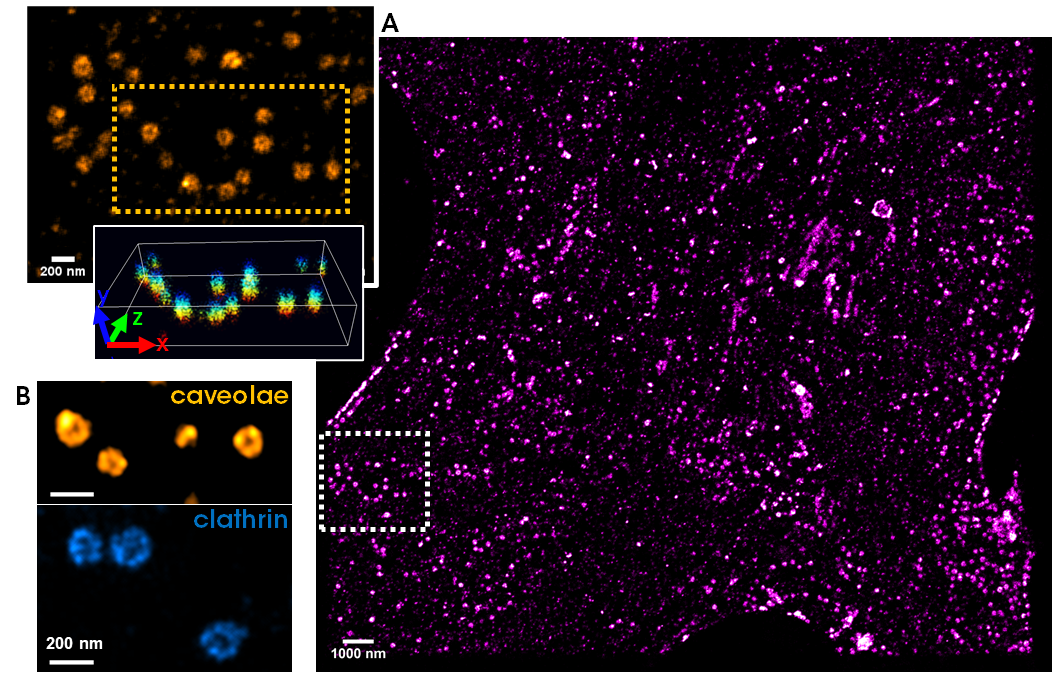
[3] Friedl, Karoline, et al. "Robust and fast multicolor Single Molecule Localization Microscopy using spectral separation and demixing." BioRxiv (2023): 2023-01.
[4] Baschieri, Francesco, et al. "Fibroblasts generate topographical cues that steer cancer cell migration." Science Advances 9.33 (2023): eade2120.
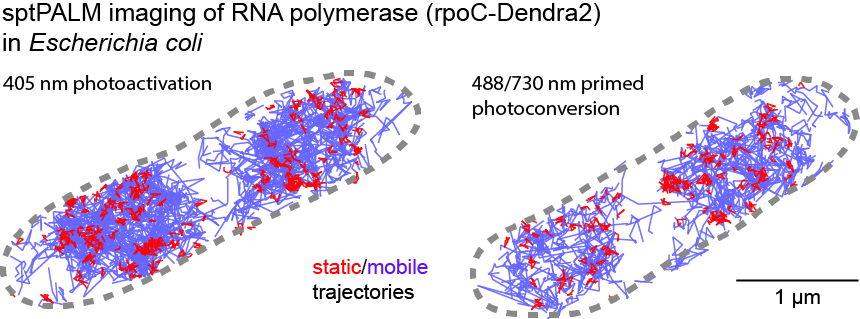
[5] Wessel, Aimee K., et al. "Escherichia coli SPFH membrane microdomain proteins HflKC contribute to aminoglycoside and oxidative stress tolerance." Microbiology Spectrum 11.4 (2023): e01767-23.
[6] Liu, Wei, et al. "Mitofusin-2 regulates leukocyte adhesion and β2 integrin activation." Journal of leukocyte biology 111.4 (2022): 771-791.
[7] Rajbanshi, Binita, et al. "Localization, induction, and cellular effects of tau phosphorylated at threonine 217." Alzheimer's & Dementia (2023).
[8] Pagliuca, M., et al. "38P Single molecule localization microscopy for extracellular vesicles detection in cancer." Annals of Oncology 33 (2022): S1395-S1396.
[9] Robaszkiewicz, A., and K. Gronkowska. "36P EP300 as an epigenetic target in p53 wild-type tumors treated with cisplatin." Annals of Oncology 33 (2022): S1395.
[10] He, Jin, et al. "Heterozygous Seryl‐tRNA Synthetase 1 Variants Cause Charcot–Marie–Tooth Disease." Annals of Neurology (2022).
[11] Gazzola, Morgan, et al. "Microtubules self-repair in living cells." Current Biology (2022).
[12] Liu, Wei, et al. "Mitofusin‐2 regulates leukocyte adhesion and β2 integrin activation." Journal of Leukocyte Biology 111.4 (2022): 771-791.
[13] Portes, Marion, et al. "Nanoscale architecture and coordination of actin cores within the sealing zone of human osteoclasts." Elife 11 (2022): e75610.
[14] Wessel, Aimee K., et al. "Escherichia coli membrane microdomain SPFH protein HflC interacts with YajC and contributes to aminoglycoside and oxidative stress tolerance." bioRxiv (2022).
[15] Radhakrishnan, A. V., et al. "Single-Protein Tracking to Study Protein Interactions During Integrin-Based Migration." The Integrin Interactome. Humana, New York, NY, (2021). 85-113.
[16] Jouchet, Pierre, et al. "Nanometric axial localization of single fluorescent molecules with modulated excitation." Nature Photonics (2021): 1-8.
[17]Orré, Thomas, et al. "Molecular motion and tridimensional nanoscale localization of kindlin control integrin activation in focal adhesions." Nature Communications 12.1 (2021): 3104.
[18] Pernier, Julien, et al. "Myosin 1b flattens and prunes branched actin filaments." Journal of cell science 133.18 (2020).
[19] Jimenez, Angélique, Karoline Friedl, and Christophe Leterrier. "About samples, giving examples: optimized single molecule localization microscopy." Methods 174 (2020): 100-114.
[20] Mau, Adrien, et al. "Fast scanned widefield scheme provides tunable and uniform illumination for optimized SMLM on large fields of view." bioRxiv (2020).
[21] Cabriel, Clément, et al. "Combining 3D single molecule localization strategies for reproducible bioimaging." Nature communications 10.1 (2019): 1980.
[22] Woodhams, Stephen G., et al. "Cell type–specific super-resolution imaging reveals an increase in calcium-permeable AMPA receptors at spinal peptidergic terminals as an anatomical correlate of inflammatory pain." Pain 160.11 (2019): 2641-2650.
[23] Belkahla, Hanen, et al. "Carbon dots, a powerful non-toxic support for bioimaging by fluorescence nanoscopy and eradication of bacteria by photothermia." Nanoscale Advances (2019).
[24] Denis, Kevin, et al. "Targeting Type IV pili as an antivirulence strategy against invasive meningococcal disease." Nature microbiology 4.6 (2019): 972.
[25] Szabo, Quentin, et al. "TADs are 3D structural units of higher-order chromosome organization in Drosophila." Science advances 4.2 (2018): eaar8082.
[26] Boudjemaa, Rym, et al. "Impact of bacterial membrane fatty acid composition on the failure of daptomycin to kill Staphylococcus aureus." Antimicrobial agents and chemotherapy 62.7 (2018): e00023-18.
[27] Culley, Siân, et al. "Quantitative mapping and minimization of super-resolution optical imaging artifacts." Nature methods 15.4 (2018): 263.
[28] Berger, Stephen L., et al. "Localized myosin II activity regulates assembly and plasticity of the axon initial segment." Neuron 97.3 (2018): 555-570.
[29] Cabriel, Clément, et al. "Aberration-accounting calibration for 3D single-molecule localization microscopy." Optics letters 43.2 (2018): 174-177.
[30] Bouissou, Anaïs, et al. "Podosome force generation machinery: a local balance between protrusion at the core and traction at the ring." ACS nano 11.4 (2017): 4028-4040.
[31] Sellés, Julien, et al. "Nuclear pore complex plasticity during developmental process as revealed by super-resolution microscopy." Scientific reports 7.1 (2017): 14732.
[32] Bourg, Nicolas, et al. "Direct optical nanoscopy with axially localized detection." Nature Photonics 9.9 (2015): 587.